Abstract
Background
Mutations in GNAS drive pancreatic tumorigenesis and frequently occur in intraductal papillary mucinous neoplasm (IPMN); however, their value as a therapeutic target is yet to be determined. This study aimed at evaluating the involvement of mutant GNAS in tumor aggressiveness in established pancreatic cancer.
Methods
CRISPR/Cas9-mediated GNAS R201H silencing was performed using human primary IPMN-associated pancreatic cancer cells. The role of oncogenic GNAS in tumor maintenance was evaluated by conducting cell culture and xenograft experiments, and western blotting and transcriptome analyses were performed to uncover GNAS-driven signatures.
Results
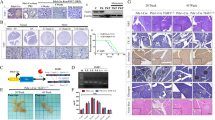
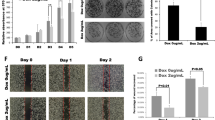
Xenografts of GNAS wild-type cells were characterized by a higher Ki-67 labeling index relative to GNAS-mutant cells. Phenotypic alterations in the GNAS wild-type tumors resulted in a significant reduction in mucin production accompanied by solid with massive stromal components. Transcriptional profiling suggested an apparent conflict of mutant GNAS with KRAS signaling. A significantly higher Notch intercellular domain (NICD) was observed in the nuclear fraction of GNAS wild-type cells. Meanwhile, inhibition of protein kinase A (PKA) induced NICD in GNAS-mutant IPMN cells, suggesting that NOTCH signaling is negatively regulated by the GNAS-PKA pathway. GNAS wild-type cells were characterized by a significant invasive property relative to GNAS-mutant cells, which was mediated through the NOTCH regulatory pathway.
Conclusions
Oncogenic GNAS induces mucin production, not only via MUC2 but also via MUC5AC/B, which may enlarge cystic lesions in the pancreas. The mutation may also limit tumor aggressiveness by attenuating NOTCH signaling; therefore, such tumor-suppressing effects must be considered when therapeutically inhibiting the GNAS pathway.





Similar content being viewed by others
References
Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Basturk O, Hong SM, Wood LD, et al. A revised classification system and recommendations from the Baltimore consensus meeting for neoplastic precursor lesions in the pancreas. Am J Surg Pathol. 2015;39:1730–41.
Ren B, Liu X, Suriawinata AA. Pancreatic ductal adenocarcinoma and its precursor lesions: histopathology, cytopathology, and molecular pathology. Am J Pathol. 2019;189:9–21.
Ryan DP, Hong TS, Bardeesy N. Pancreatic adenocarcinoma. N Engl J Med. 2014;371:1039–49.
Patra KC, Bardeesy N, Mizukami Y. Diversity of precursor lesions for pancreatic cancer: the genetics and biology of intraductal papillary mucinous neoplasm. Clin Transl Gastroenterol. 2017;8:e86.
Watanabe K, Nakamura T, Onodera S, et al. A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors. Tumour Biol. 2020;42:1010428320962588.
Drelon C, Berthon A, Sahut-Barnola I, et al. PKA inhibits WNT signalling in adrenal cortex zonation and prevents malignant tumour development. Nat Commun. 2016;7:12751.
Xing F, Luan Y, Cai J, et al. The anti-Warburg effect elicited by the cAMP-PGC1alpha pathway drives differentiation of glioblastoma cells into astrocytes. Cell Rep. 2017;18:468–81.
Furukawa T, Kuboki Y, Tanji E, et al. Whole-exome sequencing uncovers frequent GNAS mutations in intraductal papillary mucinous neoplasms of the pancreas. Sci Rep. 2011;1:161.
Wu J, Matthaei H, Maitra A, et al. Recurrent GNAS mutations define an unexpected pathway for pancreatic cyst development. Sci Transl Med. 2011;3:92ra66.
Omori Y, Ono Y, Kobayashi T, et al. How does intestinal-type intraductal papillary mucinous neoplasm emerge? CDX2 plays a critical role in the process of intestinal differentiation and progression. Virchows Arch. 2020;477:21–31.
Yamada M, Sekine S, Ogawa R, et al. Frequent activating GNAS mutations in villous adenoma of the colorectum. J Pathol. 2012;228:113–8.
Nault JC, Fabre M, Couchy G, et al. GNAS-activating mutations define a rare subgroup of inflammatory liver tumors characterized by STAT3 activation. J Hepatol. 2012;56:184–91.
Ritterhouse LL, Vivero M, Mino-Kenudson M, et al. GNAS mutations in primary mucinous and non-mucinous lung adenocarcinomas. Mod Pathol. 2017;30:1720–7.
Taki K, Ohmuraya M, Tanji E, et al. GNAS(R201H) and Kras(G12D) cooperate to promote murine pancreatic tumorigenesis recapitulating human intraductal papillary mucinous neoplasm. Oncogene. 2016;35:2407–12.
Ideno N, Yamaguchi H, Ghosh B, et al. GNAS(R201C) induces pancreatic cystic neoplasms in mice that express activated kras by inhibiting YAP1 signaling. Gastroenterology. 2018;155:1593-1607 e12.
Patra KC, Kato Y, Mizukami Y, et al. Mutant GNAS drives pancreatic tumourigenesis by inducing PKA-mediated SIK suppression and reprogramming lipid metabolism. Nat Cell Biol. 2018;20:811–22.
He X, Zhang L, Chen Y, et al. The G protein alpha subunit Galphas is a tumor suppressor in Sonic hedgehog-driven medulloblastoma. Nat Med. 2014;20:1035–42.
Iglesias-Bartolome R, Torres D, Marone R, et al. Inactivation of a Galpha(s)-PKA tumour suppressor pathway in skin stem cells initiates basal-cell carcinogenesis. Nat Cell Biol. 2015;17:793–803.
Zauber P, Marotta SP, Sabbath-Solitare M. GNAS gene mutation may be present only transiently during colorectal tumorigenesis. Int J Mol Epidemiol Genet. 2016;7:24–31.
Felsenstein M, Noe M, Masica DL, et al. IPMNs with co-occurring invasive cancers: neighbours but not always relatives. Gut. 2018;67:1652–62.
Omori Y, Ono Y, Tanino M, et al. Pathways of progression from intraductal papillary mucinous neoplasm to pancreatic ductal adenocarcinoma based on molecular features. Gastroenterology. 2019;156:647-61 e2.
Noe M, Niknafs N, Fischer CG, et al. Genomic characterization of malignant progression in neoplastic pancreatic cysts. Nat Commun. 2020;11:4085.
Kamiyama H, Kamiyama M, Hong SM, et al. In vivo and in vitro propagation of intraductal papillary mucinous neoplasms. Lab Invest. 2010;90:665–73.
Ran FA, Hsu PD, Wright J, et al. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308.
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545–50.
Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9:671–5.
Liberzon A, Birger C, Thorvaldsdottir H, et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1:417–25.
De La OJ, Emerson LL, Goodman JL, et al. Notch and Kras reprogram pancreatic acinar cells to ductal intraepithelial neoplasia. Proc Natl Acad Sci U S A. 2008;105:18907–12.
Thomas MM, Zhang Y, Mathew E, et al. Epithelial Notch signaling is a limiting step for pancreatic carcinogenesis. BMC Cancer. 2014;14:862.
Komatsu H, Tanji E, Sakata N, et al. A GNAS mutation found in pancreatic intraductal papillary mucinous neoplasms induces drastic alterations of gene expression profiles with upregulation of mucin genes. PLoS ONE. 2014;9:e87875.
Zhang H, Kong Q, Wang J, et al. Complex roles of cAMP-PKA-CREB signaling in cancer. Exp Hematol Oncol. 2020;9:32.
Pattabiraman DR, Bierie B, Kober KI, et al. Activation of PKA leads to mesenchymal-to-epithelial transition and loss of tumor-initiating ability. Science. 2016;351:aad3680.
Pan Y, Wang C, Wang B. Phosphorylation of Gli2 by protein kinase A is required for Gli2 processing and degradation and the Sonic hedgehog-regulated mouse development. Dev Biol. 2009;326:177–89.
Djouder N, Tuerk RD, Suter M, et al. PKA phosphorylates and inactivates AMPKalpha to promote efficient lipolysis. EMBO J. 2010;29:469–81.
McGill MA, McGlade CJ. Mammalian numb proteins promote Notch1 receptor ubiquitination and degradation of the Notch1 intracellular domain. J Biol Chem. 2003;278:23196–203.
Lahiry M, Kumar S, Hari K, et al. AMPK-Fyn signaling promotes Notch1 stability to potentiate hypoxia-induced breast cancer stemness and drug resistance. BioRxiv. 2020. https://doi.org/10.2139/ssrn.3586992.
Greer RL, Staley BK, Liou A, et al. Numb regulates acinar cell dedifferentiation and survival during pancreatic damage and acinar-to-ductal metaplasia. Gastroenterology. 2013;145:1088-97.e8.
Inaguma S, Kasai K, Ikeda H. GLI1 facilitates the migration and invasion of pancreatic cancer cells through MUC5AC-mediated attenuation of E-cadherin. Oncogene. 2011;30:714–23.
Valque H, Gouyer V, Gottrand F, et al. MUC5B leads to aggressive behavior of breast cancer MCF7 cells. PLoS ONE. 2012;7:e46699.
Leir SH, Harris A. MUC6 mucin expression inhibits tumor cell invasion. Exp Cell Res. 2011;317:2408–19.
Yang KS, Ciprani D, O’Shea A, et al. Extracellular vesicle analysis allows for identification of invasive IPMN. Gastroenterology. 2021;160:1345–5811.
Huang Y, Nahar S, Nakagawa A, et al. Regulation of GLI underlies a role for BET bromodomains in pancreatic cancer growth and the tumor microenvironment. Clin Cancer Res. 2016;22:4259–70.
Wilson CH, McIntyre RE, Arends MJ, et al. The activating mutation R201C in GNAS promotes intestinal tumourigenesis in Apc(Min/+) mice through activation of Wnt and ERK1/2 MAPK pathways. Oncogene. 2010;29:4567–75.
Nomura R, Saito T, Mitomi H, et al. GNAS mutation as an alternative mechanism of activation of the Wnt/beta-catenin signaling pathway in gastric adenocarcinoma of the fundic gland type. Hum Pathol. 2014;45:2488–96.
O’Hayre M, Vazquez-Prado J, Kufareva I, et al. The emerging mutational landscape of G proteins and G-protein-coupled receptors in cancer. Nat Rev Cancer. 2013;13:412–24.
Coles GL, Cristea S, Webber JT, et al. Unbiased proteomic profiling uncovers a targetable GNAS/PKA/PP2A axis in small cell lung cancer stem cells. Cancer Cell. 2020;38:129-43.e7.
Zimmerman NP, Roy I, Hauser AD, et al. Cyclic AMP regulates the migration and invasion potential of human pancreatic cancer cells. Mol Carcinog. 2015;54:203–15.
Wehbe N, Slika H, Mesmar J, et al. The role of Epac in cancer progression. Int J Mol Sci. 2020;21:6489.
Almahariq M, Chao C, Mei FC, et al. Pharmacological inhibition and genetic knockdown of exchange protein directly activated by cAMP 1 reduce pancreatic cancer metastasis in vivo. Mol Pharmacol. 2015;87:142–9.
Avila JL, Kissil JL. Notch signaling in pancreatic cancer: oncogene or tumor suppressor? Trends Mol Med. 2013;19:320–7.
Acknowledgements
We would like to thank Rika Kakisaka and Atsuko Nishikawa (Sapporo Higashi Tokushukai Hospital) for performing IHC quantification of the xenografts; Ayumu Sugitani (Sapporo Higashi Tokushukai Hospital) for supporting statistical analyses; and Eiko Aoyanagi (Sapporo Higashi Tokushukai Hospital) for tissue sample preparation. We also thank Krushna C. Patra (Department of Cancer Biology at University of Cincinnati) for critical reading of the manuscript.
Funding
This work was supported by Pancreas Research Foundation of Japan and Grants-in-Aid for Regional R&D Proposal-Based Program from Northern Advancement Center for Science & Technology of Hokkaido in Japan (to H. Kawabata) and by JSPS KAKENHI Grant Number 20K17009 (to H. Kawabata), 20K07671 (to Y.O.), 20K09070 (to H. Karasaki) and 20H03655 (to Y.M.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
YO and YM receive funding from Hitachi High-Tech Corporation. The other authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kawabata, H., Ono, Y., Tamamura, N. et al. Mutant GNAS limits tumor aggressiveness in established pancreatic cancer via antagonizing the KRAS-pathway. J Gastroenterol 57, 208–220 (2022). https://doi.org/10.1007/s00535-021-01846-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-021-01846-4