Abstract
GNAS mutations have been described in mucinous and non-mucinous epithelial neoplasms of the appendix, pancreas, and colon, with hotspot GNAS mutations found in up to two-thirds of pancreatic intraductal papillary mucinous neoplasms. Additionally, many GNAS-mutated tumors have concurrent mutations in the Ras/Raf pathway. The clinicopathologic features of GNAS-mutated lung carcinomas, however, have not yet been characterized. Primary lung carcinomas from Brigham and Women's Hospital (n=1282) or Massachusetts General Hospital (n=1070) were genotyped on a targeted massively parallel sequencing panel of oncogenes and tumor suppressor genes including GNAS. Clinical and pathological features were reviewed, and TTF-1 immunohistochemistry was performed when material was available. Nineteen lung adenocarcinomas with hotspot GNAS mutations were identified (19/2352, 0.8%) including 14 at codon 201 and 5 at codon 227. GNAS-mutated lung adenocarcinomas occurred predominantly in female patients (16/19, 84%). Ten (10) were classified as invasive mucinous adenocarcinomas (IMA), and nine (9) were non-mucinous adenocarcinomas. All IMAs had GNAS codon 201 mutations and concurrent Ras/Raf pathway mutations (9 KRAS, 1 BRAF). No tumors with GNAS codon 227 mutations had mucinous histological features. 86% of GNAS-mutated non-mucinous adenocarcinomas (6/7) were positive for TTF-1 immunohistochemistry, while only 25% of GNAS-mutated IMAs (1/4) were positive for TTF-1. Patients with GNAS-mutated non-mucinous adenocarcinomas were more likely to have a history of smoking (9/9, 100%) compared to patients with GNAS-mutated IMAs (2/10, 20%) (P<0.001). Hotspot GNAS mutations can occur in primary lung adenocarcinomas. When associated with concurrent mutations in the Ras/Raf pathway, these neoplasms often present as IMAs. GNAS mutations are not specific to neoplasms of the gastrointestinal tract, and clinicopathologic correlation is necessary in GNAS-mutated adenocarcinomas in the lung to determine the primary site of origin.
Similar content being viewed by others
Main
GNAS hotspot mutations have been described in various epithelial neoplasms, including appendiceal, pancreatic, and colonic tumors, and are found in up to two-thirds of intraductal papillary mucinous neoplasms (IPMNs).1, 2, 3, 4, 5 GNAS is a proto-oncogene originally described in pituitary adenomas that encodes for the Gsα subunit of heterotrimeric G-proteins that transduce signals via the G-stimulatory pathway and increase production of cyclic AMP (cAMP).6 Hotspot mutations have been identified in codons 201 and 227, with both hotspots resulting in constitutive activation of the G-stimulatory pathway with increased cAMP production.6, 7
GNAS mutations in pancreatic cyst fluid have been shown to be a helpful biomarker in classifying pancreatic cystic lesions and are highly specific for IPMNs in that setting.1, 2 GNAS mutations have also been identified in invasive pancreatic adenocarcinomas arising from IPMNs but rarely in invasive carcinomas without associated IPMNs.1 GNAS mutations are present in ~20% of appendiceal mucinous tumors including low-grade appendiceal mucinous neoplasms and mucinous adenocarcinomas.3 Additionally, ~2% of colonic adenocarcinomas harbor hotspot GNAS mutations and have been associated with a villous morphology and right-sided location.4 Another study has found a correlation between GNAS mutations in colonic adenocarcinomas and a mucinous phenotype, with hotspot GNAS mutations seen in 20% of the mucinous colonic adenocarcinomas in the study.8 GNAS-mutated neoplasms in the colon and pancreas frequently harbor concurrent mutations in KRAS, and are seen concurrently in ~90% of GNAS-mutated colon adenocarcinomas and in 50% of GNAS-mutated IPMNs.1, 4
Five to ten percent of lung adenocarcinomas are invasive mucinous adenocarcinomas (IMA).9, 10, 11 Mucinous neoplasms of the lung can be diagnostically challenging, as these tumors can be morphologically and immunohistochemically identical to metastatic mucinous tumors of the gastrointestinal tract.12, 13, 14, 15, 16, 17, 18 Primary IMA of the lung have distinctive molecular alterations, including frequent pathway-activating mutations in KRAS, occasional loss of function mutations in NKX2-1, and rare rearrangements in ALK and NRG1.19, 20, 21, 22, 23, 24 Although GNAS mutations have been described in association with mucinous and non-mucinous tumors in multiple organs, pathway-activating GNAS mutations have yet to be characterized in lung carcinomas. Assessing whether GNAS mutations can also be associated with primary lung adenocarcinomas could be of potential use to determine primary site of origin of mucinous tumors in the lung. In this study, we interrogate 2352 lung carcinomas subjected to targeted somatic tumor sequencing panels from two institutions for the presence of activating GNAS mutations and evaluate their associated clinicopathologic features.
Materials and methods
Case Selection
This study was approved by the institutional review boards at Dana Farber Cancer Institute and Massachusetts General Hospital and included patients with lung carcinoma undergoing a targeted next generation sequencing assay performed on tumor tissue at the Center for Advanced Molecular Diagnostics (n=1282) (Department of Pathology, Brigham and Women's Hospital) and the Center for Integrated Diagnostics (n=1070) (Department of Pathology, Massachusetts General Hospital). Only cases that were determined to be a primary lung carcinoma based on clinical history, radiologic findings, and pathologic features were included.
Clinicopathologic Features
The following features were recorded in the cases examined: sex, age at diagnosis, clinical presentation, radiographic findings, smoking history, and clinical stage. Histological slides were reviewed, and classification was performed according to the International Association for the Study of Lung Cancer recommendations.25 IMA was defined by goblet or columnar cell morphology with abundant intracytoplasmic mucin in greater than 90% of the tumor cells.
Immunohistochemistry
Primary antibody for TTF-1 was applied to five micron glass slides of formalin fixed, paraffin-embedded tumor tissue (clone 8G6G3/1 Dako, Inc., 1:300 dilution in citrate buffer, pressure cooker). Dako EnVision+ (Agilent Technologies, Santa Clara, CA) was used for signal detection. Strong nuclear immunohistochemical staining for TTF-1 in tumor cells was considered positive.
Targeted Tumor Genomic Sequencing (‘OncoPanel’) at the Center for Advanced Molecular Diagnostics
OncoPanel tumor profiling was performed as previously described.26 Indexed sequencing libraries were prepared from a 50-ng sonically sheared DNA sample using Illumina TruSeq LT reagents (Illumina, Inc., San Diego, CA). A custom RNA bait set by Agilent SureSelect (Agilent Technologies, Santa Clara, CA) was used to enrich libraries for exons and select introns of 282 genes implicated in cancer biology by using solution-based hybrid capture (list of genes and transcripts, Supplementary Table 1). Massively parallel sequencing was performed using Illumina HiSeq2500 (Illumina, Inc.) with 100 × 100 paired-end reads to achieve a mean target coverage greater than 50 ×. Sample reads were analyzed using a custom pipeline including Picard, BWA, and GATK (Broad Institute, Cambridge, MA) for sorting and alignment. Mutation analysis for single-nucleotide variants was performed using MuTect version 1 0.27200 (Broad Institute, Cambridge, MA). Insertions and deletions were called using Indelocator. Variants were filtered to exclude known germline variants in the Single-Nucleotide Polymorphism database by MuTect and variants that occur at a population frequency >0.1% in the Exome Sequencing Project database.
Targeted Tumor Genomic Sequencing at the Center for Integrated Diagnostics
Mutation hotspots and exons from 39 genes were targeted using Anchored Multiplex PCR.27 Total nucleic acid containing total RNA and genomic DNA were extracted from formalin-fixed, paraffin-embedded tissue by using the Agencourt FormaPure Kit (Beckman Coulter, Indianapolis, IN, https://www.beckmancoulter.com) before undergoing end-repair, adenylation, and adapter ligation. Two hemi-nested PCR reactions were then performed using one primer specific to a sequence in the gene of interest and one specific to a universal sequence in the adapter, for each PCR. This library was then sequenced on the Illumina MiSeq (Illumina, San Diego, CA) and 2 × 150 base paired-end reads were aligned to the human genome hg19 reference sequence with the BWA-MEM software package (version bwa-0.7.5a).28 Single-nucleotide variants were detected with MuTect and indels with a custom indel-detection algorithm.29 In regions with sufficient read coverage, this approach is validated to detect indels and single-nucleotide variants with allelic frequencies as low as 5%. The complete list of the genes and exons interrogated is listed in Supplementary Table 2.
Statistical Analysis
Categorical data was analyzed using Fisher's exact test. Statistical analysis was two-sided, and a P-value of <0.05 was considered statistically significant.
Results
Clinicopathologic Features of GNAS-Mutated Lung Carcinomas
The combined cohort of lung carcinomas (n=2352) included the following histotypes: 79% adenocarcinoma, 12% squamous cell carcinoma, 7% non-small cell lung carcinomas not otherwise specified, 1% large cell neuroendocrine carcinoma, and 1% adenosquamous carcinoma. Hotspot GNAS mutations involving codons 201 and 227 were identified in 11 of 1282 lung carcinomas (0.9%) at the Center for Advanced Molecular Diagnostics and in 8 of 1070 lung carcinomas (0.7%) at the Center for Integrated Diagnostics (total 19/2352, 0.8%), including 5 at codon 227 and 14 at codon 201 (Table 1). Abdominal and pelvic imaging study results were available for review in 18/19 patients with 15/18 (83%) having completely negative findings, 1 with multiple liver and bony metastases (case 2), 1 with multiple liver, pancreas, and adrenal gland lesions thought to represent metastases (case 9), and 1 with a 1 cm cystic lesion in the pancreas (case 17). Follow-up MRI imaging was performed on the cystic pancreatic lesion, which revealed a side-branch IPMN. This side-branch IPMN was considered to be indolent in nature and unrelated to the patient's lung tumor. In all 19 GNAS-positive cases, the patients were clinically determined to have primary lung tumors.
Patients with GNAS-mutated lung carcinomas had a median age of 66 (range 24–78). GNAS-mutated lung carcinomas occurred predominantly in female patients (16/19, 84%). Ten of 19 GNAS-mutated lung carcinomas (53%) were classified as IMAs, whereas the remaining 9 tumors were adenocarcinomas without mucinous histological features (Figure 1). No cases met the criteria for colloid adenocarcinoma in the material that was available for review. TTF-1 immunohistochemistry was performed when tissue was available (n=11) and was positive in 6/7 non-mucinous adenocarcinomas (86%) and 1/4 IMA (25%) (P=0.09, Figure 2). Patients with GNAS-mutated non-mucinous adenocarcinomas were more likely to have a history of smoking (9/9, 100%) than those with IMAs (2/10, 20%) (P<0.001) (Figure 3).
GNAS-mutated lung carcinoma histology. (a) Case 3 with GNAS p.Gln227Leu, non-mucinous adenocarcinoma histology with papillary and micropapillary architecture. (b) Case 4 with GNAS p.Gln227Leu, non-mucinous adenocarcinoma histology with acinar architecture. (c) Case 12 with GNAS p.Arg201Cys and KRAS p.Gly12Asp, invasive mucinous adenocarcinoma. (d) Case 17 with GNAS p.Arg201Cys and KRAS p.Gly13Asp, invasive mucinous adenocarcinoma.
Clinical presentation of IMAs included upper respiratory infection with cough in 2 cases, incidentally identified lung nodule in 4 cases, abdominal pain in 1 case, follow-up imaging for a history of esophageal adenocarcinoma in 1 case, and chronic cough with shortness of breath in 1 case (Supplementary Table 3). Radiographic presentation included multiple bilateral lung nodules in 4 cases, cavitary or cystic nodules in 3 cases, and groundglass opacities in 6 cases (Supplementary Table 3). Sixty percent of IMAs (6/10) were stage I at the time of diagnosis compared to 22% of non-mucinous adenocarcinomas (2/9); however, this correlation did not reach statistical significance (P=0.17).
GNAS codon 227 Mutated Lung Carcinomas Have Non-Mucinous Adenocarcinoma Histology
Five neoplasms had GNAS p.Gln227 mutations (4 p.Gln227Leu, 1 p.Gln227His) (Table 1). All of the GNAS p.Gln227 variants were transversion mutations (A>T (n=4), G>T (n=1)) and had non-mucinous adenocarcinoma histology (Figures 1a and b). The morphologies among these 5 cases included a mix of acinar, papillary, micropapillary, solid, and lepidic patterns. One case had a concurrent KRAS mutation (p.Gly12Cys), but did not show mucinous histologic features.
GNAS codon 201 Mutated Lung Carcinomas with Concurrent Raf/Ras Pathway Mutations are Associated with Invasive Mucinous Adenocarcinoma Histology
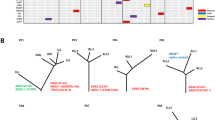
Fourteen neoplasms had GNAS p.Arg201 mutations (6 p.Arg201Cys, 6 p.Arg201His, 1 p.Arg201Leu, 1 p.Arg201Ser). Additionally, a subset of these tumors (12/14, 86%) had concurrent mutations in the Ras/Raf pathway (9 KRAS, 1 HRAS, 2 BRAF) (Table 1). All IMA had GNAS p.Arg201 mutations and concurrent Ras/Raf pathway mutations (9 KRAS, 1 BRAF) (Figures 1c and d). A large percentage (78%) of KRAS mutations in IMAs with concurrent GNAS p.Arg201 mutations were G>A transitions. Similarly, all of the GNAS p.Arg201 variants seen in IMAs were also transitions (G>A (n=4) or C>T (n=6)). Case 7 had GNAS p.Arg201Leu and HRAS p.Gln61Arg mutations, and while there was only a cytologic specimen available for review, no mucinous features were present. Additionally, Case 8 had GNAS p.Arg201His and BRAF p.Gly469Ala mutations but did not have histological features of IMA.
Additional Genomic Alterations in GNAS-Mutated Lung Adenocarcinomas
Recurrent mutations in TP53 and CDKN2A were identified in the cohort of 19 GNAS-mutated lung adenocarcinomas. TP53 mutations were seen in 3 of 5 (60%) adenocarcinomas with GNAS p.Gln227 mutations and 4 of 14 (29%) adenocarcinomas with GNAS p.Arg201 mutations. CDKN2A mutations were seen in 1 of 5 (20%) adenocarcinomas with GNAS p.Gln227 mutations and 2 of 14 (14%) adenocarcinomas with GNAS p.Arg201 mutations. TP53 variants included nonsense, splice site, and missense mutations, including TP53 p.Tyr220Cys, p.Arg249Gly, and p.Arg273Leu hotspot mutations. TP53 mutations were more likely to be seen in non-mucinous GNAS-mutated adenocarcinomas than in GNAS-mutated IMAs, 67 vs 10%, respectively (P=0.02). CDKN2A variants were frameshift or splice site variants, implying loss of tumor suppressor function (Table 1).
Discussion
GNAS mutations are most commonly associated with gastrointestinal neoplasms, including pancreatic IPMNs and invasive adenocarcinomas arising from IPMNs, colonic adenocarcinomas and mucinous appendiceal tumors.1, 2, 3, 4, 5, 8 Our study demonstrates that GNAS mutations are present in a small subset (0.8%) of primary lung carcinomas. As the genomic profiling of lung adenocarcinomas becomes more clinically utilized, it is important to recognize that GNAS variants can be identified in primary lung adenocarcinomas and do not necessarily indicate metastasis from a gastrointestinal site. Fifteen patients in this series underwent abdominal imaging without evidence of extrapulmonary disease. Although this study cannot absolutely exclude that these neoplasms may represent metastases from undiagnosed gastrointestinal sites, we consider this possibility unlikely given the patients' clinical evaluations and the known pathogenesis of pancreatic and other gastrointestinal mucinous neoplasms.
Mutations in both GNAS hotspots, p.Arg201 and p.Gly227, are seen in lung carcinomas. Approximately half of GNAS-mutated lung carcinomas have IMA histology, all of which have concurrent GNAS p.Arg201 and Ras/Raf pathway mutations. In contrast, GNAS p.Gly227 mutations are not associated with IMAs. The pathobiology behind this association is uncertain, as both p.Arg201 and p.Gly227 GNAS hotspot mutations have similar mechanisms of action, inhibiting GTPase activity, and thereby functioning as a dominant oncogene by constitutively activating Gs.6
Patients with IMA histology with concurrent GNAS p.Arg201 and Ras/raf pathway alterations are significantly less likely to have a history of smoking than those patients with GNAS p.Gln227 mutations and non-mucinous histology. Although KRAS mutations in lung carcinoma,30, 31 particularly KRAS transversion mutations (G>T,G>C) have a strong association with smoking,32, 33 7 of the 9 IMAs with concurrent GNAS p.Arg201 and KRAS mutations in our cohort had KRAS G>A transition mutations, which is more commonly seen in non-smokers. The presence of these ‘gastrointestinal-type’ KRAS mutations (G>A transitions) has been previously described in primary lung IMAs and should not be taken as evidence that a tumor is a metastasis from a gastrointestinal primary site.34 Additionally, it should be emphasized that the presence of a GNAS mutation cannot be used as a tool to distinguish primary lung adenocarcinomas from gastrointestinal or pancreaticobiliary metastases, as GNAS-mutated tumors from each of these sites have overlapping morphologic, immunophenotypic, as well as molecular findings. Therefore, careful clinicopathologic correlation is required in these scenarios to properly identify primary site of disease.
IMAs comprise a subset of lung adenocarcinomas with distinctive morphological and clinical features. IMAs are becoming an increasingly heterogeneous molecular group. In addition to common oncogenic mutations in KRAS, IMAs may infrequently harbor ALK and NRG1 rearrangements.24 Recurrent NKX2-1 (also known as TTF1) mutations have been identified in conjunction with KRAS mutations, and have been shown to promote pathogenesis and loss of TTF-1 expression in mucinous lung cancer.23 This study shows an association between IMA histology and mutations involving KRAS and GNAS p.Arg201, suggesting a novel mechanism by which cooperative KRAS and GNAS mutations promote primary lung carcinogenesis toward a mucinous or gastrointestinal phenotype. Understanding the association between secondary mutations in KRAS-mutated lung adenocarcinoma and morphological phenotype requires additional experimentation and in vivo models.
The clinical and biological significance of GNAS mutations in lung carcinoma is currently unknown. Some data suggests that patients with pseudomyxoma peritonei and GNAS mutations have a significantly shorter progression free survival than patients without GNAS mutations.35 Additionally, the same study has found that GNAS-mutated colorectal carcinomas have an aggressive clinical course. In contrast, GNAS-mutated IPMN-associated carcinomas follow a more indolent course than those with GNAS mutations36
Currently, therapies directly targeting GNAS mutations are not available. However, based on preclinical evidence, tumors with GNAS mutations may respond to inhibitors of the MAPK pathway.37 More data is needed to determine whether GNAS mutations in lung carcinoma portend a particular clinical outcome or will be responsive to a specific targeted therapy. Since IMAs are heterogeneous on the molecular level, and the histological features of GNAS-mutated lung adenocarcinomas are not distinctive, broad testing of lung adenocarcinomas should be employed to identify such cases, and will become increasingly necessary should a therapeutic agent become available.
Although GNAS mutations are present in an infrequent subset of lung adenocarcinomas, the overall incidence of lung cancer is high, and the prevalence of GNAS-mutated lung adenocarcinomas is expected to be significant. This group of neoplasms is important to recognize and correctly classify the primary site based on clinicopathologic features, which can be challenging in the setting of IMA histology and negative TTF-1 immunohistochemistry. Further studies will be needed to determine their clinical course, biologic behavior, and whether they will be amenable to targeted therapies.
References
Wu J, Matthaei H, Maitra A et al. Recurrent GNAS mutations define an unexpected pathway for pancreatic cyst development. Sci Transl Med 2011;3:92ra66.
Springer S, Wang Y, Dal Molin M et al. A combination of molecular markers and clinical features improve the classification of pancreatic cysts. Gastroenterology 2015;149:1501–1510.
Hara K, Saito T, Hayashi T et al. A mutation spectrum that includes GNAS, KRAS and TP53 may be shared by mucinous neoplasms of the appendix. Pathol Res Pract 2015;211:657–664.
Fecteau RE, Lutterbaugh J, Markowitz SD et al. GNAS mutations identify a set of right-sided, RAS mutant, villous colon cancers. PLoS ONE 2014;9:e87966.
Kadayifci A, Atar M, Wang JL et al. Value of adding GNAS testing to pancreatic cyst fluid KRAS and carcinoembryonic antigen analysis for the diagnosis of intraductal papillary mucinous neoplasms. Dig Endosc 2017;29:111–117.
Landis CA, Masters SB, Spada A et al. GTPase inhibiting mutations activate the alpha chain of Gs and stimulate adenylyl cyclase in human pituitary tumours. Nature 1989;340:692–696.
Lyons J, Landis CA, Harsh G et al. Two G protein oncogenes in human endocrine tumors. Science 1990;249:655–659.
Stachler MD, Rinehart E, Lindeman N et al. Novel molecular insights from routine genotyping of colorectal carcinomas. Hum Pathol 2015;46:507–513.
Yoshizawa A, Motoi N, Riely GJ et al. Impact of proposed IASLC/ATS/ERS classification of lung adenocarcinoma: prognostic subgroups and implications for further revision of staging based on analysis of 514 stage I cases. Mod Pathol 2011;24:653–664.
Lee HY, Cha MJ, Lee KS et al. Prognosis in resected invasive mucinous adenocarcinomas of the lung: related factors and comparison with resected nonmucinous adenocarcinomas. J Thorac Oncol 2016;11:1064–1073.
Kadota K, Yeh YC, D'Angelo SP et al. Associations between mutations and histologic patterns of mucin in lung adenocarcinoma: invasive mucinous pattern and extracellular mucin are associated with KRAS mutation. Am J Surg Pathol 2014;38:1118–1127.
Goldstein NS, Thomas M . Mucinous and nonmucinous bronchioloalveolar adenocarcinomas have distinct staining patterns with thyroid transcription factor and cytokeratin 20 antibodies. Am J Clin Pathol 2001;116:319–325.
Saad RS, Cho P, Silverman JF et al. Usefulness of Cdx2 in separating mucinous bronchioloalveolar adenocarcinoma of the lung from metastatic mucinous colorectal adenocarcinoma. Am J Clin Pathol 2004;122:421–427.
Tsuta K, Ishii G, Nitadori J et al. Comparison of the immunophenotypes of signet-ring cell carcinoma, solid adenocarcinoma with mucin production, and mucinous bronchioloalveolar carcinoma of the lung characterized by the presence of cytoplasmic mucin. J Pathol 2006;209:78–87.
Stenhouse G, Fyfe N, King G et al. Thyroid transcription factor 1 in pulmonary adenocarcinoma. J Clin Pathol 2004;57:383–387.
Wislez M, Antoine M, Baudrin L et al. Non-mucinous and mucinous subtypes of adenocarcinoma with bronchioloalveolar carcinoma features differ by biomarker expression and in the response to gefitinib. Lung Cancer 2010;68:185–191.
Wu J, Chu PG, Jiang Z et al. Napsin A expression in primary mucin-producing adenocarcinomas of the lung: an immunohistochemical study. Am J Clin Pathol 2013;139:160–166.
Shah RN, Badve S, Papreddy K et al. Expression of cytokeratin 20 in mucinous bronchioloalveolar carcinoma. Hum Pathol 2002;33:915–920.
Finberg KE, Sequist LV, Joshi VA et al. Mucinous differentiation correlates with absence of EGFR mutation and presence of KRAS mutation in lung adenocarcinomas with bronchioloalveolar features. J Mol Diagn 2007;9:320–326.
Sakuma Y, Matsukuma S, Yoshihara M et al. Distinctive evaluation of nonmucinous and mucinous subtypes of bronchioloalveolar carcinomas in EGFR and K-ras gene-mutation analyses for Japanese lung adenocarcinomas: confirmation of the correlations with histologic subtypes and gene mutations. Am J Clin Pathol 2007;128:100–108.
Kakegawa S, Shimizu K, Sugano M et al. Clinicopathological features of lung adenocarcinoma with KRAS mutations. Cancer 2011;117:4257–4266.
Ichinokawa H, Ishii G, Nagai K et al. Distinct clinicopathologic characteristics of lung mucinous adenocarcinoma with KRAS mutation. Hum Pathol 2013;44:2636–2642.
Hwang DH, Sholl LM, Rojas-Rudilla V et al. KRAS and NKX2-1 mutations in invasive mucinous adenocarcinoma of the lung. J Thorac Oncol 2016;11:496–503.
Fernandez-Cuesta L, Plenker D, Osada H et al. CD74-NRG1 fusions in lung adenocarcinoma. Cancer Discov 2014;4:415–422.
Travis WD, Brambilla E, Noguchi M et al. International association for the study of lung cancer/american thoracic society/european respiratory society international multidisciplinary classification of lung adenocarcinoma. J Thorac Oncol 2011;6:244–285.
Sholl LM, Do K, Shivdasani P et al. Institutional implementation of clinical tumor profiling on an unselected cancer population. JCI Insight 2016;1:e87062.
Zheng Z, Liebers M, Zhelyazkova B et al. Anchored multiplex PCR for targeted next-generation sequencing. Nat Med 2014;20:1479–1484.
Li H, Durbin R . Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009;25:1754–1760.
Cibulskis K, Lawrence MS, Carter SL et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 2013;31:213–219.
Buttitta F, Barassi F, Fresu G et al. Mutational analysis of the HER2 gene in lung tumors from Caucasian patients: mutations are mainly present in adenocarcinomas with bronchioloalveolar features. Int J Cancer 2006;119:2586–2591.
Suzuki M, Shigematsu H, Iizasa T et al. Exclusive mutation in epidermal growth factor receptor gene, HER-2, and KRAS, and synchronous methylation of nonsmall cell lung cancer. Cancer 2006;106:2200–2207.
Ahrendt SA, Decker PA, Alawi EA et al. Cigarette smoking is strongly associated with mutation of the K-ras gene in patients with primary adenocarcinoma of the lung. Cancer 2001;92:1525–1530.
Riely GJ, Kris MG, Rosenbaum D et al. Frequency and distinctive spectrum of KRAS mutations in never smokers with lung adenocarcinoma. Clin Cancer Res 2008;14:5731–5734.
Shim HS, Kenudson M, Zheng Z et al. Unique genetic and survival characteristics of invasive mucinous adenocarcinoma of the lung. J Thorac Oncol 2015;10:1156–1162.
Pietrantonio F, Berenato R, Maggi C et al. GNAS mutations as prognostic biomarker in patients with relapsed peritoneal pseudomyxoma receiving metronomic capecitabine and bevacizumab: a clinical and translational study. J Transl Med 2016;14:125.
Tan MC, Basturk O, Brannon AR et al. GNAS and KRAS mutations define separate progression pathways in intraductal papillary mucinous neoplasm-associated carcinoma. J Am Coll Surg 2015;220: 845–854 e1.
Wilson CH, McIntyre RE, Arends MJ et al. The activating mutation R201C in GNAS promotes intestinal tumourigenesis in Apc(Min/+) mice through activation of Wnt and ERK1/2 MAPK pathways. Oncogene 2010;29:4567–4575.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
AJI is a stockholder and scientific advisory board member of ArcherDx and is a consultant for Roche, Pfizer, DebioPharm, Chugai, and Constellation Pharmaceuticals. MM-K is a consultant for Merrimack pharmaceuticals, H3 biomedicine, ACD, and Roche. Authors LLR, MV, VN, LMS, and FD have no disclosures.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Ritterhouse, L., Vivero, M., Mino-Kenudson, M. et al. GNAS mutations in primary mucinous and non-mucinous lung adenocarcinomas. Mod Pathol 30, 1720–1727 (2017). https://doi.org/10.1038/modpathol.2017.88
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.88
This article is cited by
-
Oncogene addiction to GNAS in GNASR201 mutant tumors
Oncogene (2022)
-
Mutant GNAS limits tumor aggressiveness in established pancreatic cancer via antagonizing the KRAS-pathway
Journal of Gastroenterology (2022)
-
Molecular subtypes based on DNA methylation predict prognosis in lung squamous cell carcinoma
BMC Cancer (2021)
-
Diffuse expression of MUC6 defines a distinct clinicopathological subset of pulmonary invasive mucinous adenocarcinoma
Modern Pathology (2021)
-
Elevated expression of GNAS promotes breast cancer cell proliferation and migration via the PI3K/AKT/Snail1/E-cadherin axis
Clinical and Translational Oncology (2019)